Overview of research
Our team studies the molecular mechanisms responsible for preventing alterations in numbers and structure of chromosomes. Our current efforts are targeted at understanding how changes in chromatin structure are regulated by the cell cycle machinery during mitosis. We also study a human cancer-predisposition disease –the Nijmegen breakage syndrome– characterized by defects in cell cycle checkpoint regulation. Cells from patients with this inherited disease have a very severe deficiency in the detection and repair of DNA damage, often resulting in altered chromosome structure
As a whole, our research program integrates fundamental aspects of cell biology such as how cell signaling by proteases and protein kinases regulates cell cycle progression and chromatin structure in dividing cells.
Specific research projects are described under the section “experimental approaches” below.
Experimental Approaches
We address our biological questions using one of the very best model organisms available for the analysis of eukaryotic cell cycle regulation, the budding yeast Saccharomyces cerevisiae. We combine the use of this model organism with cutting edge technologies and experimental approaches, including proteomics (mass spectrometry, phosphorylation analyses, chromatin complex purification), cell biology (live cell microscopy, cell synchronization) and molecular genetic analyses.
SPECIFIC PROJECTS
Regulation of chromosome structure during cell division
Click for DetailsSignaling cascades regulating cell division: How cells control exit from mitosis?
Click for DetailsRegulation of genomic integrity by SMC complexes:The cases of the Nijmegen breakage syndrome and Smc5/6 complex
Click for Details1 – Regulation of chromosome structure during cell division
The total length of naked DNA in the human genome is ~2 meters when placed head-to-toe. How can this fit into a cell smaller than 0.1 mm?
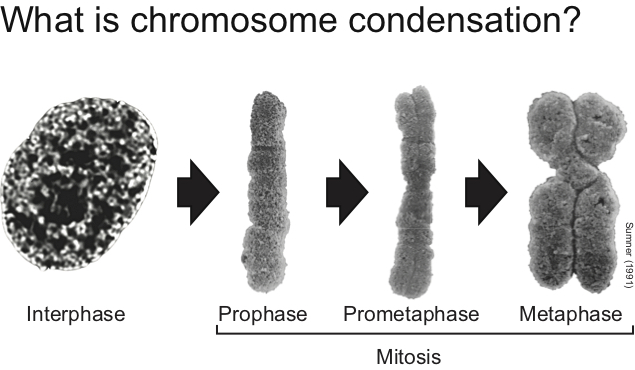
This is a basic question of great importance in modern biology. Eukaryotic cells have resolved this issue by folding DNA into chromatin, thereby allowing some degree of DNA compaction. However, the levels of chromatin compaction achieved in normal resting cells (i.e., GI or G2) is insufficient to allow high fidelity chromosome transmission during mitosis. So, what happens with chromatin structure when cells divide their chromosomes during cell division? This is the specific question we aim to answer.
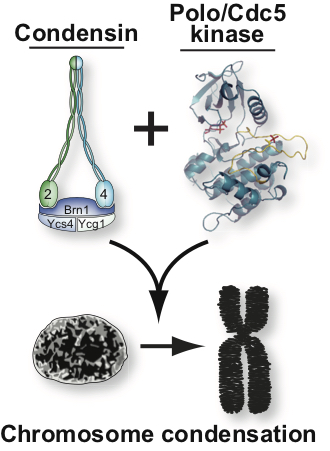
Genetic studies have revealed that a key factor for the regulation of chromatin structure in mitosis is condensin, a pentameric protein complex used by the cell to assemble fully condensed chromosomes. Despite much progress in recent years regarding condensin’s role in chromosome condensation, relatively little is known at present on how this essential complex may regulate chromosome structure.
Our work is focussed on addressing the following major questions on chromosome condensation:
- How is chromatin folded in the highly-condensed sister-chromatids of mitotic chromosomes?
- How does condensin promote efficient formation of condensed chromatin during mitosis?
- How is condensin activated and inactivated during mitosis?
2 – Signaling cascades regulating cell division: How cells control exit from mitosis?
The cell cycle can be viewed as alternating phases of high and low cyclin-dependent kinase (CDK) activity. It is well known that cells require high CDK-cyclin B activity to enter mitosis whereas they need to inactivate CDK-cyclin B to exit mitosis and return to a G1 state. How does a cell decide when is the right time to exit mitosis and complete cell division?
Inactivation of CDK activity at the end of mitosis requires the integration of many cellular cues to make sure that completion –or “exit”– from mitosis occurs exactly at the right time. We study the signaling cascades responsible for inducing exit from mitosis. There are two main signal transduction pathways responsible for ending mitosis: The mitotic exit network (MEN) and the Cdc14 early anaphase release network (FEAR network).
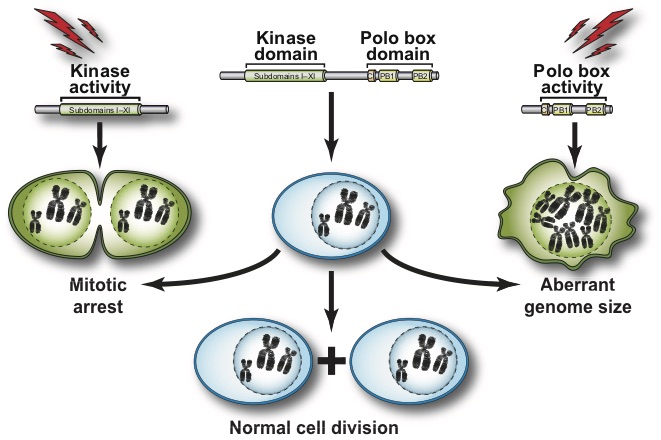
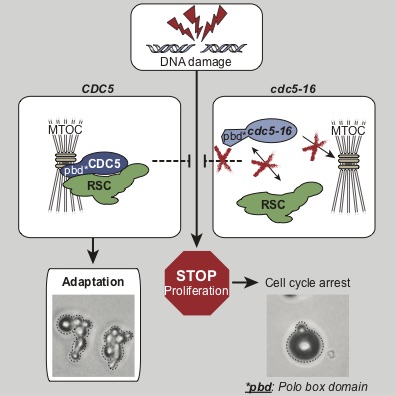
We focus our analyses of cell cycle progression and mitotic exit an evolutionarily conserved kinase, the Polo kinase Cdc5 (see images on the right for a summary of its role in cell cycle and checkpoint responses). This kinase is a shared component of the MEN and the FEAR networks and misregulation of the human homolog of this kinase is implicated in many different forms of cancers.
Our work is aimed at addressing the following major questions on the specific mode of action of this cancer-associated kinase:
- How is this kinase activated during mitosis?
- What are the substrates of this kinase during cell division?
- What are the contributions of the kinase domain and the polo-box domain (PDB) of Cdc5 to its cellular functions?
3 – Regulation of genomic integrity by SMC complexes:
The cases of the Nijmegen breakage syndrome and Smc5/6 complex
Cancer cells can modify their genome by inducing structural defects in their genome, such as the fusion of DNA sequences from unrelated chromosomes. With this specific project, our laboratory is interested in understanding the molecular mechanisms responsible for preventing alterations in the basic structure of chromosomes.
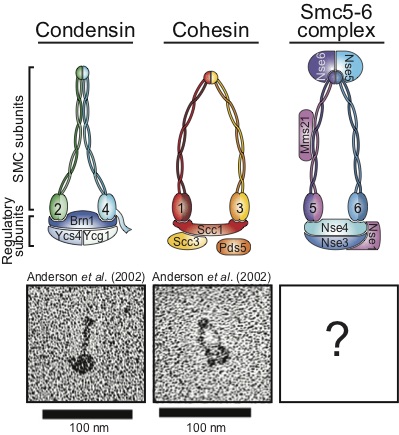
We know that a family of proteins known as the Structural Maintenance of Chromosome (SMC) proteins plays an essential role in the maintenance of genome stability. This family of proteins is characterized by a unique structural organization. The four conserved eukaryotic SMC complexes play a wide variety molecular functions including key roles in DNA repair and checkpoint signaling. In spite of recent advances in our knowledge, we still only have a partial mechanistic understanding of how SMC complexes promote DNA repair and genome integrity.
Our current efforts are targeted at understanding a human cancer-predisposition disease -the Nijmegen breakage syndrome (“NBS”)- characterized by defects in cell cycle checkpoint regulation in response to DNA damaging agents. Cells from patients with this inherited disease have a deficiency in a member of the Rad50 SMC complex, the Nbs1/Xrs2 protein.
NBS cells have a very severe deficiency in the detection and repair of DNA damage, often resulting in altered chromosome structure. Understanding the molecular defects underlying this disease represent an invaluable opportunity to find key cellular players involved in cancer development.
Finally, we also study a related complex involved the detection of DNA damage, the Smc5/6 complex, in order to understand what may be the common functions of the Rad50 and Smc5/6 complexes in the detection and signaling of DNA damage.